LUYOR-3109高强度紫外催化光源促销
LUYOR-3109紫外光源采用了9颗365nm大功率led,安装有二次光学透镜,输出紫外线强度高,...
2024-08-08作者:上海路阳生物技术时间:2021-12-27 11:13:16浏览2002 次
In summary, on the basis of transcriptomics analysis of two types of R. sphaeroides and enzymatic activity verification, we identified Prsp_7571 as a native highly active promoter. Through random sequence mutation and the use of an RFP visualization screening system, we constructed a Prsp_7571-derived synthetic promoter library. Finally, a selected strong synthetic promoter was used to significantly improved the Q10 biosynthetic efficiency in a high-producing industrial strain. These well-characterized promoters should be highly useful in current synthetic biology platforms for refactoring the biosynthetic pathway in R. sphaeroides-derived MCFs.
中国农业大学生物学院农业技术实验室使用LUYOR-3415RG便携式荧光蛋白激发光源照射的文献近期发表。文献地址:
https://doi.org/10.1016/j.ibmb.2021.103703
A cytorhabdovirus-based expression vector in Nilaparvata lugens, Laodelphax striatellus, and Sogatella furcifera
Abstract
The brown planthopper (BPH, Nilaparvata lugens), the small brown planthopper (SBPH, Laodelphax striatellus), and the white-backed planthopper (WBPH, Sogatella furcifera) are problematic insect pests and cause severe yield losses through phloem sap-sucking and virus transmission. Barley yellow striate mosaic virus (BYSMV), a plant cytorhabdovirus, has been developed as versatile expression platforms in SBPHs and cereal plants. However, bio-safe overexpression vectors based on recombinant BYSMV (rBYSMV) remain to be developed and applied to the three kinds of planthoppers. Here, we found that rBYSMV was able to infect SBPHs, BPHs and WBPHs through microinjection with crude extracts from rBYSMV-infected barley leaves. To ensure bio-safety of the rBYSMV vectors, we generated an rBYSMV mutant by deleting the accessory protein P3, a putative viral movement protein. As expected, the resulting mutant abolished viral systemic infection in barley plants but had no effects on BYSMV infectivity in insect vectors. Subsequently, we used the modified rBYSMV vector to overexpress iron transport peptide (ITP) in the three kinds of planthoppers and revealed the potential functions of ITP. Overall, our results provide bio-safe overexpression platforms to facilitate functional genomics studies of planthoppers.
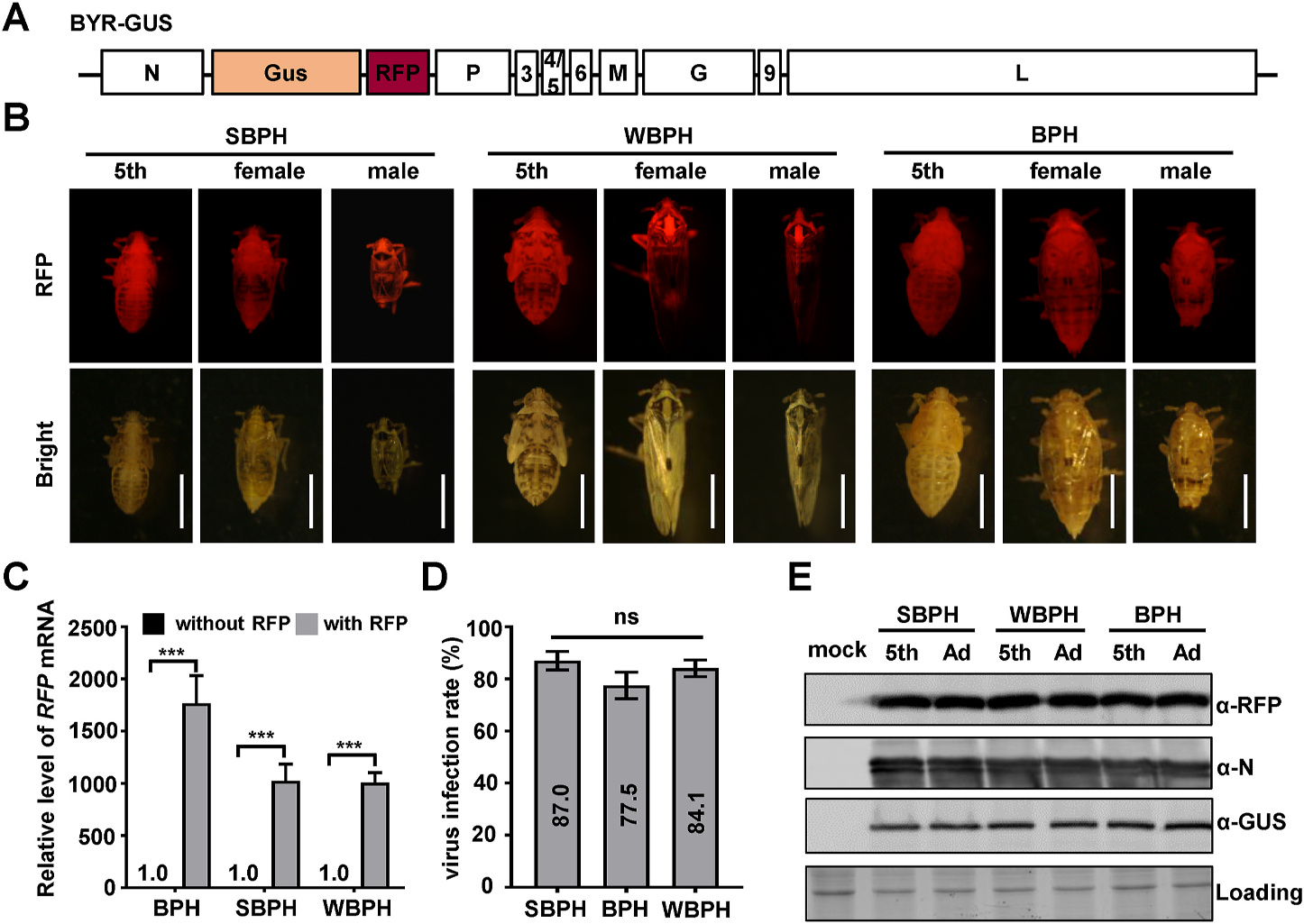
上图的荧光图片是在LUYOR-3415RG照射下拍摄的
Introduction
In East and Southeast Asia, three species of planthoppers, including the brown planthopper (BPH), the small brown planthopper (SBPH), and the white-backed planthopper (WBPH) are devastating agricultural pests that lead to drying of plants called “hopper burn” (Backus et al., 2005; Cheng, 2015; He et al., 2019). In addition, these planthoppers served as transmission vectors of many economically important plant viruses (Cao et al., 2018; Dader et al., 2017; Zheng et al., 2014; Zhou et al., 2013). Pioneering functional genomic studies have significantly improved our understanding of planthopper outbreaks, insect-virus interactions and pest management (Poelchau et al., 2016). In the last two decades, RNAi and CRISPR/Cas9 have been widely used in studying gene functions of various insects. RNAi is a powerful approach for sequence-specific gene silencing, in which dsRNA corresponding to genes of interest are microinjected into insects directly or delivered along with artificial diet (Basnet and Kamble, 2018; Chen et al., 2010; Jain et al., 2020; Liu et al., 2010; Zhu and Palli, 2020). Moreover, the genomes of BPH, SBPH, and WBPH have been sequenced and are available for designing specific dsRNA templates (Wang et al., 2017; Xue et al., 2014; Zhu et al., 2017). Thus, various genes involved in reproduction, wing polyphenism, development, and survival have been identified using RNAi (Hu et al., 2019; Liu et al., 2020; Qiu et al., 2016; Shen et al., 2021; Xu et al., 2015; Yang et al., 2021).
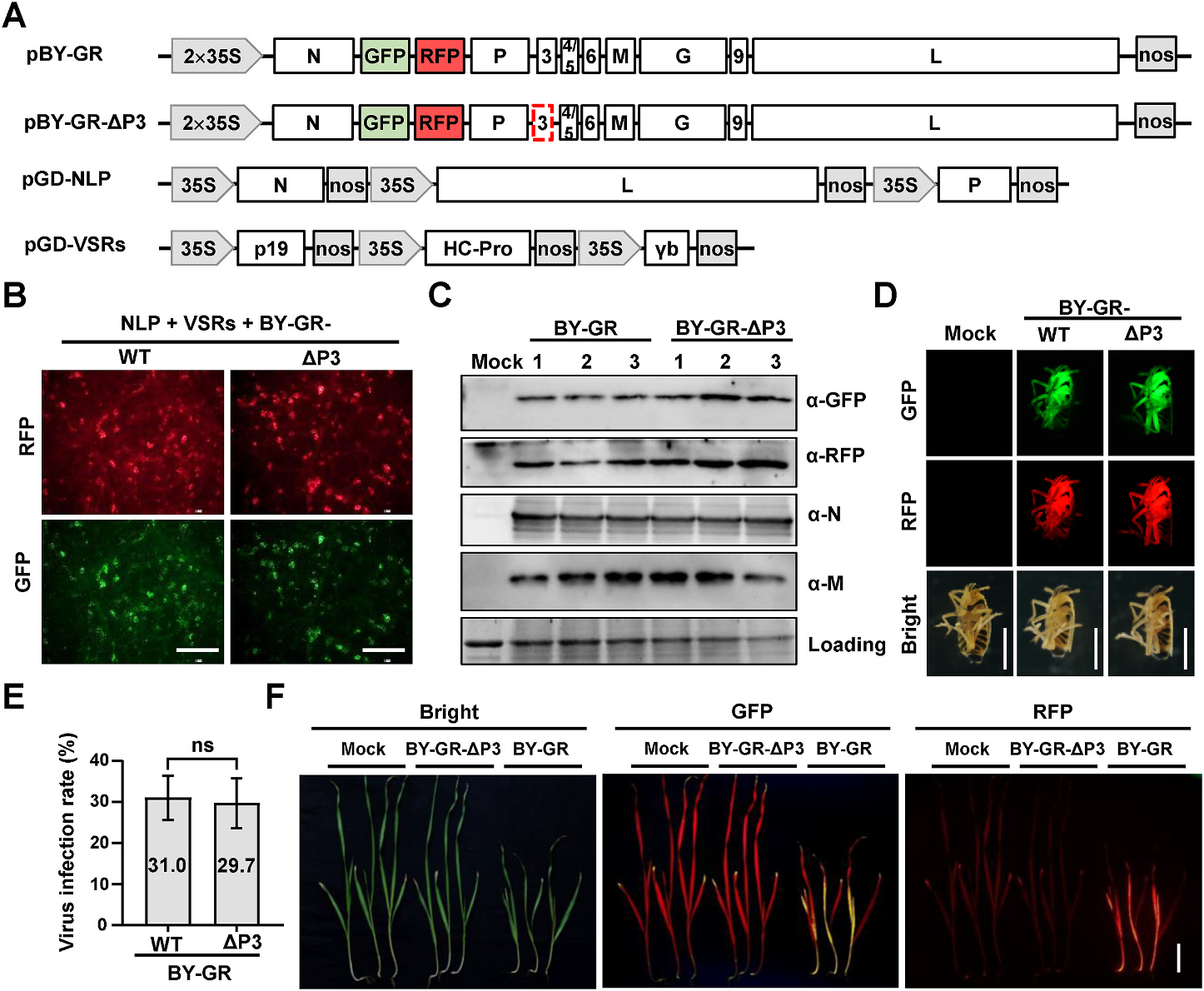
Recently, the CRISPR/Cas9 systems have been applied to generation of stable and heritable mutants for genomic studies (Jinek et al., 2012; Wang et al., 2016). The first application of CRISPR/Cas9 technology in insects resulted in a 4.6 kb deleted mutant in a yellow locus of Drosophila melanogaster (Gratz et al., 2013). Subsequently, the CRISPR/Cas9 system had been developed in several insect orders including Hymenoptera, Diptera, Lepidoptera, Coleoptera, Hemiptera, and Orthoptera (Bi et al., 2016; Taning et al., 2017; You et al., 2019; Zhang et al., 2015). The first attempt of genome editing in planthopper was achieved in BPHs, but the hatchability of injected-eggs was less than 5% (Xue et al., 2018). Afterwards, an optimized strategy was established with hatchability up to 10.5% (Zhao et al., 2019). In addition to RNAi-induced gene knockdown and CRISPR/cas9-mediated gene knockout, gene overexpression is an important strategy in fundamental genetic analysis of insects (Fraser, 2012). Barley yellow striate mosaic virus (BYSMV), a cytorhabdovirus, has been engineered as a versatile expression platform in SBPHs (Gao et al., 2019). However, the virus-mediated overexpression vectors have not been reported in BPHs and WBPHs. In this regard, versatile overexpression approaches are crucial for gene functional studies in planthoppers.
The iron transport peptide (ITP) and its alternatively spliced variant ITP-like (ITPL) belong to the crustacean hyperglycemic hormone family (Dircksen et al., 2001). ITP, but not ITPL, harbors a canonical C-terminal amidation signal (-GKR, -GKK, or -GRK) (Wang et al., 2000), and both are widely conserved among various insect species (Dircksen, 2009; Webster et al., 2012). ITP is detected in brains, whereas ITPL localizes in the peripheral nervous systems and other tissues, implying their specialized physiological functions (Dai et al., 2007; Endo et al., 2000). ITP/ITPL are first identified to play roles in ion and fluid transport across the ileum in Schistocerca gregaria (Audsley et al., 1992; Meredith et al., 1996; Pfeifer et al., 1999). Subsequently, ITPs have been shown to be critical for ecdysis in Manduca sexta and moulting, development, and oviposition in Tribolium castaneum (Begum et al., 2009; Drexler et al., 2007). Interestingly, ITP is also a functional clock neuropeptide in Drosophila melanogaster and played a critical role in water homeostasis (Galikova et al., 2018; Hermann-Luibl et al., 2014). In Bombyx mori, mutations in receptor of ITP resulted in a prolongation of the larval stage and failure in wing expansion of moths (Sun et al., 2020). In Nilaparvata lugens, individuals injected with dsNilluITP/ITPL showed crumpled wings and cuticle melanism after eclosion (Yu et al., 2016). Nonetheless, whether overexpression of ITP affects developments of planthoppers remains to be investigated.
Barley yellow striate mosaic virus (BYSMV), a negative-stranded virus with the genome size of 12.7 kb, is a member of the genus Cytorhabdovirus and transmitted by SBPHs in a propagative manner (Cao et al., 2018). The BYSMV genome encodes 10 proteins in the order 3′-N-P-P3-P4/P5–P6-M-G-P9-L-5′ (Yan et al., 2015). The N, P, L, M, and G proteins are structural proteins mainly involved in virus replication, packaging and recognition with receptors (Jackson et al., 2005). In addition, plant rhabdoviruses encode accessory proteins at the position between P and M genes or position between G and L genes (Jackson et al., 2005). The P3 proteins of several rhabdoviruses have been identified as cell-to-cell movement proteins in plants (Huang et al., 2005; Mann et al., 2016 Zhou et al., 2019). We first developed a mini-replicon system of BYSMV in Nicotiana benthamiana (N. benthamiana) leaves (Fang et al., 2019). Subsequently, we rescued the full length of recombinant BYSMV vectors as delivery and expression platforms in SBPHs and cereals (Gao et al., 2019). Thereafter, crude extract from BY-RFP-infected barley was co-injected with dsRNA into SBPHs to investigate the functions of targeted genes in BYSMV infection (Gao et al., 2020; Zhang et al., 2020). Nonetheless, a bio-safe BYSMV-based vector is still required for genomic studies of planthoppers.
In this study, we found that BYSMV was able to infect SBPHs, WBPHs and BPHs by microinjection. We further showed that the BYSMV P3 protein was essential for viral systemic infection in host plant but not for propagation in insect vectors. Accordingly, a bio-safe rBYSMV mutant without accessory protein P3 was generated to investigate the potential functions of ITP in WBPHs, BPHs, and SBPHs.